Ligand Dose Response for Different Affinity Synthetic Receptors on an Engineered Biomaterial Surface
ABSTRACT
Stem cells have the unique ability to self-renew by creating new stem cells and differentiate into many cell types. By leveraging these abilities, this project seeks to create a platform capable of consistently generating biomimetic tissues by better controlling differentiation processes using synthetic receptors. We utilized the synthetic receptor synNotch and engineered it into stem cells in addition to developing a compatible biomaterial surface. Our biomaterial immobilizes activating ligands which are then presented to stem cells resulting in activation. To optimize this platform, we studied two synNotch receptors, the high-affinity LaG16 and low-affinity LaG17 anti-GFP receptors, by assessing their activation thresholds in the presence of immobilized GFP. Six concentrations of soluble GFP ranging from 0-200 nM were used with either a biomaterial surface or a control surface. The high-affinity LaG16 synNotch receptor was able to activate at the lowest concentration of 2.5 nM GFP on the engineered surface while the cells on the control surface remained inactivated until 50 nM GFP. These results suggest that 2.5 nM to 5 nM is the ideal activation range for LaG16 synNotch. Preliminary data for the low affinity LaG17 synNotch receptor showed faint activation at 2.5 nM that continued to increase with GFP concentration.
INTRODUCTION.
Stem cells have two unique characteristics: the ability to self-renew and the ability to differentiate into many cell types (1). The three main types of stem cells are somatic/adult, embryonic, and induced pluripotent stem cells. Somatic stem cells are found in adults and are very limited in terms of what they can differentiate into. Embryonic stem cells are present in embryos and can differentiate into any cell in the body. Embryonic stem cells have been used in the past to treat neurodegenerative diseases (1). Induced pluripotent stem cells are reprogrammed somatic cells that have the same pluripotency as embryonic stem cells.
Stem cell differentiation can mimic development, in which adoption of tissue-specific cell fates is preceded by the establishment of the three germ layers. The three germ layers are the mesoderm which gives rise to muscle and bone cells, the ectoderm which gives rise to skin and nervous system cells, and the endoderm which gives rise to respiratory and digestive tract cells (2). Stem cells also have the ability to self-renew by dividing into identical cells. Stem cells have become a topic of interest in medicine due to their regenerative capabilities (1). One way to further leverage this attribute is through the use of synthetic receptors, such as synNotch, to influence differentiation into desired cell types for the creation of biomimetic tissues.
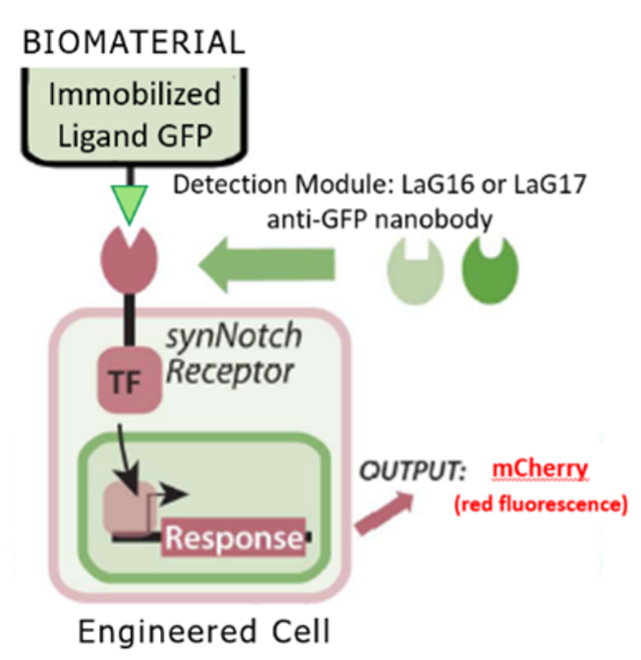
The synNotch synthetic signaling platform responds to immobilized ligands resulting in downstream target gene expression (4). The synNotch platform is based off the native juxtracrine Notch/Delta signaling channel, meaning that synNotch requires an immobilized ligand for activation. This feature leads to highly localized receptor activation. SynNotch is also a programmable synthetic receptor in which the user can define the intracellular and extracellular domains. The intracellular domain refers to the transcription factor which causes gene expression in the nucleus and the extracellular domain refers to part of the receptor that detects the activating ligand. When presented with an activating ligand, an intramembrane cleavage occurs in the synthetic receptor which results in the release of a transcription factor leading to downstream expression of target genes (Figure 1).
Figure 1. The synNotch receptor (4). Activation starts when the detection module captures an immobilized ligand. In this experiment, the synNotch receptors used contained either a LaG16 or LaG17 GFP responsive detection module. The transcription factor is then released via intramembrane cleavage. The transcription factor then goes to the nucleus where it triggers expression of target genes, such as the fluorescent reporter mCherry. Figure adapted from Lee et al. (3) Made in Biorender
The purpose of this project was to engineer a biomaterial surface that could then be used to test two different receptors with varying affinities for green fluorescent protein (GFP). The biomaterial that was engineered in this project consisted of two adhesion peptides and one ligand capturing peptide. The adhesion peptides were GAG-Binding Peptide (GBP) (5) and cycloRGD (CRGD) (6) which are Glycosaminoglycan (GAG) and integrin binding respectively. The ligand capturing peptide was GFP Trap (3) which was responsible for immobilizing green fluorescent protein ligands and presenting them to the cells.
This experiment utilized an immobilized GFP ligand to activate an anti-GFP detection module which resulted in the expression of an mCherry protein reporter. The two cell lines engineered through isothermal assembly contained one of two variations of synNotch which were LaG16 and Lag17. LaG17 was prepared using a lentiviral vector while sleeping beauty was used for the LaG16 cell line. The main difference between these two synthetic receptors is their affinity for GFP: LaG16 has a higher affinity (Kd = 0.7nM) and requires less GFP to activate it while LaG17 is lower affinity (Kd = 50nM) and requires more GFP to activate. By understanding the ideal activation range of different affinity synNotch receptors by dosing them with a broad range of GFP concentrations, better control over the surface dependent activation of cells can be achieved.
This platform could take advantage of digital fabrication and micropatterning to produce tissue engineering substrata or scaffolds to spatially dictate stem cell behaviors in response to global soluble cues. Such an advance could lead to improved engineered tissue substitutes or methods to produce organoids for studying development and disease. By only engineering certain cells or using specific concentrations of activating ligands, cells could be patterned to differentiate into different cell types, giving the ability to generate biomimetic tissues more consistently.
We hypothesize that the higher synNotch activation will be seen on the higher affinity receptor compared to the lower affinity receptor and the biomaterial will be required for activation at lower concentrations.
MATERIALS AND METHODS.
Cell Engineering. Through isothermal assembly, two H9 embryonic stem cell lines were engineered to express different affinity synthetic receptors. The stem cells were engineered with a mCherry transgene reporter insert and a synNotch receptor backbone. Plasmid assembly consists of 5 main steps: digestion, gel extraction, isothermal assembly, transformation, and purification. In digestion, the insert (gene of interest) and backbone (plasmid DNA) are separated from their larger plasmids via enzymes. In gel extraction, samples are run through gel electrophoresis to isolate the insert and backbone DNA. In isothermal assembly, using the Gibson Assembly method, the insert and backbone are mixed together and combine to form a plasmid through activity of an exonuclease, polymerase, and ligase enzyme mix. In transformation, the plasmid is added to bacteria to increase the amount of DNA by utilizing the fast speed at which bacteria proliferate and replicate plasmids. In purification, the bacteria are lysed to release the plasmid which is then purified from the solution. H9 stem cells were transfected with the purified plasmid. The plasmids contained a puromycin resistant gene allowing us to select the engineered cells.
Cell Maintenance. The cells were maintained in mTeSR Plus media (Stem Cell Technologies, Cat #: 100-0276). Cells were plated on a Geltrex® (Fisher Scientific, Cat # A1413302) coated 6 well in ~ 1:10 split. Routine passages were done with ReLeSR at ~70-80% confluence.
Media preparation. To prepare the stock media, stem cell maintenance media, mTeSR, was supplemented with 10 µM rock inhibitor (Tocris, Cat. #: 1254) which is a substance that promotes viability of stem cells in single-cell suspensions. A portion of the stock media was supplemented with 200 nM GFP. This 200 nM concentration of GFP was diluted with the stock media to obtain the 2.5, 5, 50, and 100 nM GFP concentrations.
Biomaterial preparation. For the engineered condition, well plates were coated with 10 µg/mL streptavidin (Thermo Scientific, Cat. #: 21125B) and incubated overnight. GBP (Genscript, custom peptide synthesis), cRGD (Fisher Scientific, Cat. #: 50-168-6291), and GFP Trap were mixed together in a 5: 2.15: 0.8 molar ratio to form the biomaterial. CRGD (integrin binding) and GBP (GAG binding) are adhesion peptides and Peg-GFP Trap is a ligand capturing peptide. To create a functionalized cell culture surface, the peptide solution (75 µL) was then added to each well of a 96-well cell culture dish and the plates were incubated for 1 hour before cell dissociation. For the control surface, well plates were coated in 75 µL of Geltrex® basement membrane.
Cell Dissociation. Cells were passaged using Accutase to create a single cell suspension. The cells were incubated for 5 minutes. The cells were then quenched with mTeSR media (1 mL). Cells were counted and split equally into six Eppendorf tubes which were spun down. The supernatant was removed and the cells were resuspended in mTeSR media with varying GFP concentrations. ~40,000 cells were plated per well in a 96 well plate. There were 3 wells per experimental condition on either the engineered surface or the control surface for a total of 36 wells.
Data processing and Statistical Analysis. Microscopy data was collected using a Leica DMi8 Microscope using phase contrast and fluorescence channels. Images were taken on day two prior to performing flow cytometry. Mean fluorescence intensity was gathered using a CellStream flow cytometer. Data was analyzed using FlowJo software to isolate the cell populations of interest. Samples were gated for a cell population, a single cell population, and an mCherry positive population. Mean mCherry fluorescence was calculated for the single cell population.
Data was analyzed using a one-way ANOVA to check for significant differences between experimental groups. Following ANOVA analysis, post hoc Tukey tests were performed to further determine the statistical difference across all experimental groups.
RESULTS.
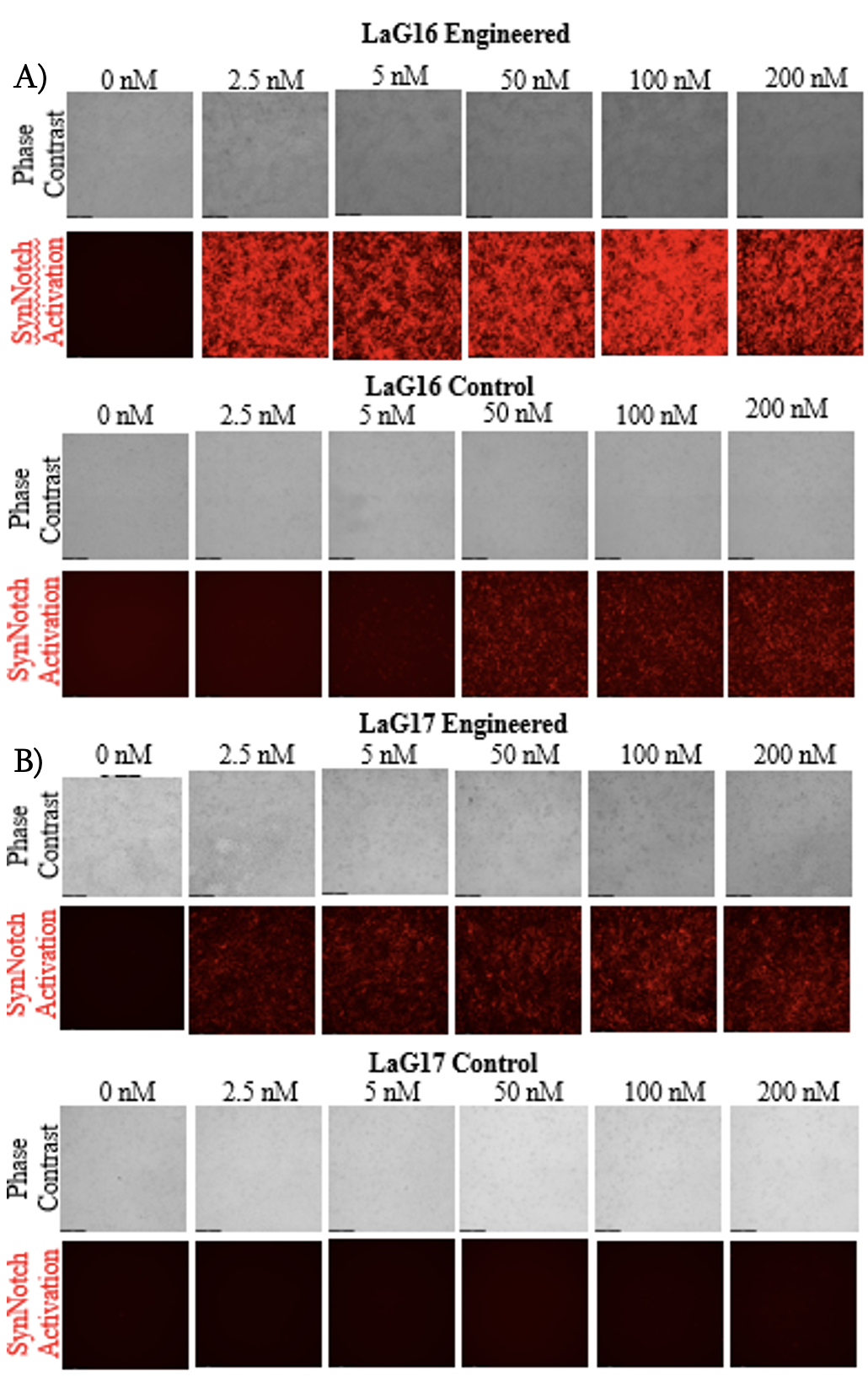
The following results were obtained from the LaG16 cell line by preforming microscopy and flow cytometry. Phase contrast and fluorescence images were collected on day two of culture (Figure 2). The cells were imaged for two main reasons: to view the adherence of cells to the substrate and to visualize the synNotch activation through mCherry expression. The microscopy revealed that cells were able to adhere to the substrate in all tested conditions. Additionally, synNotch activation was observed at GFP concentrations as low as 2.5 nM.
Figure 2. Part A is microscopy images of Lag16 activation (Engineered on top, control on bottom). Scale bar=200 µm. Part B is microscopy images of Lag17 (Engineered on top, control on bottom). The sheer difference between the engineered and control surface activation in both conditions are visualized in these images.
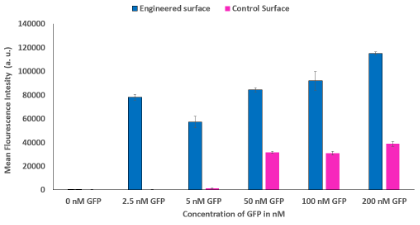
To quantify synNotch activation, flow cytometry was performed, and the mCherry fluorescence intensity data was analyzed using Flowjo. A shift in the cell population towards mCherry positive (increase in fluorescence intensity) was observed at all concentrations on the engineered surface and the higher concentrations on the control surface. The mean fluorescence intensity of each condition was calculated to compare synNotch activation thresholds (Figure 3). The flow analysis shows that there was an activation peak at a range from 2.5 nM GFP to 5 nM GFP for Lag16. Additionally, a one-way ANOVA revealed significant differences between every concentration and 0 nM on the engineered surface. Furthermore, the 2.5, 50, and 100 nM GFP groups did not show a significant difference in activation. The ANOVA also revealed a significant difference between the higher concentrations (50, 100, and 200 nM) and the lower concentrations (0, 2.5, and 5 nM) on the control surface.
Figure 3. SynNotch Activation on Engineered Surface Compared to Control Surface. Compares the mean fluorescence intensity of the engineered surface and the control surface on LaG16. Engineered surface activation starts at the lowest concentration while control surface activation is not detected until 50 nM GFP. Asterisks signify significant differences.
DISCUSSION.
The purpose of this project was to analyze the activation thresholds of two synthetic receptors. For the high affinity LaG16 receptor, the activation threshold was determined to be at 2.5 nM GFP or less. Even though additional activation can be seen past this range (Figure 3), surface independent activation of the control surface occurred at GFP concentrations above 5 nM due to an abundance of activating ligand. For the purpose of micropatterning, use of minimal ligand concentration could result in cells on an engineered surface signaling via synNotch while leaving cells on a control surface. Additionally, there is no significant difference between 0, 2.5, and 5 nM GFP on the control surface, meaning that there is negligible activation on the control surface at the lower concentrations. This supports the hypothesis that the engineered surface could be activated independently of the control surface enabling the consistent creation of micropattern tissues. Additionally, the microscopy indicated that there was strong cell adherence as well as no peeling on both the engineered and control surfaces in all conditions (Figure 2). The lack of peeling observed on both the engineered and control surfaces is consistent with the conclusion that the biomaterial surface has the same cell adherence as traditionally surfaces, such as the control in this project.
For the LaG17 synNotch receptor, the microscopy shows faint synNotch activation at the lowest GFP concentration of 2.5 nM GFP (Figure 2). As the concentration of GFP increased to 200 nM GFP, there was an increase in mCherry fluorescence intensity. This indicates that as the GFP concentration increases, so too does the synNotch activation. The LaG17 synNotch receptor showed a fainter mCherry fluorescence intensity than the LaG16 synNotch receptor at the lower GFP concentrations which is likely due to its lower affinity for GFP. While activation of the LaG17 synNotch receptor is surprising, it could be due to the different plasmid delivery method used for generating the cell line. Additionally, the microscopy data displays the similarity in cell adherence for both conditions. This further suggests that the biomaterial is just as effective at enabling cell adhesion whether synNotch is active or not. Based on these results, we can expand on micropatterning by applying different affinity cells instead of a control or engineered surface.
Results obtained from this study could lead to the creation of a platform capable of consistently generating biomimetic tissues by utiliz ing the activation thresholds of synthetic receptors when ligand presenting biomaterial surfaces are used. This idea is supported by both the increased sensitivity of the cells on the biomaterial surface at lower GFP concentrations as well as the sheer number of mCherry positive cells on the engineered surface when compared to the control surface. Future studies will seek to spatially constrain our engineered synNotch cells to develop biomimetic tissues.
CONCLUSION.
This project should be considered a preliminary study since it utilized 3 samples and larger sample sizes should be used in future studies. A limitation of this study was that the number of synNotch positive cells were not determined, meaning that the true activation range of LaG16 synNotch cannot be accurately defined. This could also explain the dip in activation observed at 5 nM GFP. For consistency, future studies should use one plasmid delivery method for both cell lines. Future studies should attempt to use the sleeping beauty plasmid delivery method for both cell lines. Future studies should focus on gathering flow analysis for the LaG17 synNotch receptor. Additionally, future studies should use the applications of this research for micropatterning to spatially constrain the cell responses to GFP ligand. In conclusion, the ideal activation range for LaG16 synNotch is 2.5 nM GFP or less. Using a higher concentration will lead to oversaturation, resulting in the loss of surface dependent activation which is needed to micropattern effectively. Additionally, the cells were able to adhere to the engineered substratum in every condition, meaning that it is an effective replacement for traditional surfaces.
ACKNOWLEDGMENTS.
I would like to thank Hannah Brien for mentoring and helping me with this project. I would also like to thank Catherine Hamann for co-mentoring me. Joanne Lee provided technical input with the PEG-GFP Trap utilized in this project. Thank you to Dr. Bunger and the SSMV for giving me this opportunity. I am very grateful to the entirety of the Brunger Lab for all their support. Experimental reagents were supplied thanks to support from NSF RECODE CBET 2033800.
REFERENCES
- W. Zakrzewski, M. Dobrzyński, M. Szymonowicz, & Z. Rybak, Stem cells: Past, present, and future. Stem Cell Research & Therapy, 10(1). (2019).
- L. Solnica-Krezel, & D. S. Sepich, Gastrulation: Making and shaping germ layers. Annual Review of Cell and Developmental Biology, 28(1), 687–717. (2012).
- Lee et al., Programmable Orthogonal Cell-Biomaterial Interaction for Regenerative Engineering. [Manuscript submitted for publication to Biomaterials]. BME Vanderbilt University. (2022).
- L. Morsut, K. T. Roybal, X. Xiong, R. M. Gordley, S. M. Coyle, M. Thomson, & W. A. Lim, Engineering customized cell sensing and response behaviors using synthetic notch receptors. Cell, 164(5), 780–791. (2016).
- P. J. Wrighton et al., Signals from the surface modulate differentiation of human pluripotent stem cells through glycosaminoglycans and integrins. Proceedings of the National Academy of Sciences of the United States of America, 111 (51) 18126-18131. (2014).
- J. R. Klim et al., A defined glycosaminoglycan-binding substratum for human pluripotent stem cells. Nature Methods, 7, 989–994. (2010).
Posted by John Lee on Tuesday, May 30, 2023 in May 2023.
Tags: Biomaterials, Pluripotent Stem Cells, Regenerative Engineering, Synthetic Receptors, Tissue Engineering